iBioSim Documentation
Introduction
iBioSim has been developed for the modeling, analysis, and design of genetic circuits. While iBioSim primarily targets models of genetic circuits, models representing metabolic networks, cell-signaling pathways, and other biological and chemical systems can also be analyzed. iBioSim also includes modeling and visualization support for multi-cellular and spatial models as well. It is capable of importing and exporting models specified using the Systems Biology Markup Language (SBML).Finally, it is one of the first tools to also support the Synthetic Biology Open Language (SBOL), an emerging standard for information exchange in synthetic biology.
IMPORTANT: this documentation is still under development. Older documentation and tutorials are found here:
- Version : v 3.0
- Created by the Myers Research Group
- GitHub Repository : https://github.com/SynBioDex/SBOLDesigner
- Issue Tracker : https://github.com/SynBioDex/SBOLDesigner/issues
- Download : JAR File
- Readme : README.md
To cite iBioSim, please use the latest paper:
Watanabe, Leandro, Tramy Nguyen, Michael Zhang, Zach Zundel, Zhen Zhang, Curtis Madsen, Nicholas Roehner, and Chris Myers. 2018. "IBioSim 3: A Tool for Model-Based Genetic Circuit Design." ACS Synthetic Biology 8 (7): 1560–1563. doi:10.1021/acssynbio.8b00078.
Installing iBioSim #back to top
Before installing iBioSim, please make sure that the following software is installed on your system.
- Java Runtime Environment (Version 1.7 or higher)
- Graphviz (optional)
To install, download the zip file for your operating system and unzip it where you want the application to reside.
To run iBioSim, execute iBioSim in the top directory.
For Windows users:
If the application font appears very small, you need to follow the next steps:
- Go to where Java is installed (usually C:\Program Files\Java\jre\bin) and right click on the java.exe file. Select Properties, click the Compatibility tab, click on "Change high DPI settings", and select "Override high DPI scaling behavior." and finally select Scaling performed by: System.
- If the previous step didn't solve the problem, try repeating the last steps for all java*.exe files (i.e: java.exe, javaw.exe javacl.exe, etc).
- If the previous step didn't solve the problem, open a Command Prompt and type "where ava". This will list if you have any other folders containing java.exe files. Find these other files, and follow the instructions in step 1) of each of them.
For Linux users:
Dependencies are resolved at runtime when iBioSim is opened
for the first time.
In order
to compile the dependencies, you need the following:
- bison
- flex
- swig
- python2
- libxml2-dev
Getting Started #back to top
Before beginning to use iBioSim, go to your preferences and change your name, email, and URI to something unique.
Preferences Screen
Creating a Project #back to top
To create a new project, select New → Project from the File menu as shown below. You will then be prompted to browse to a desired location and to give a name to the project directory. After you do this, click the new button and a new project directory will be created. To open a project, select Open Project from the File menu. You will then be prompted to browse to a project directory to open, and clicking open will open the project. You may also open a project by selecting one of your ten most recently opened projects by selecting the project name shown in the Open Recent menu within the File menu.
Main Screen
This is the main screen of iBioSim. The top bar comprises of functions such as saving and exporting, while the bottom bar holds parts available for use.
Importing from Repositories #back to top
To import a registry part from an online repository, double click on a part to view its details, click on (import registry part) then choose and filter the desired repository such as synBioHub.
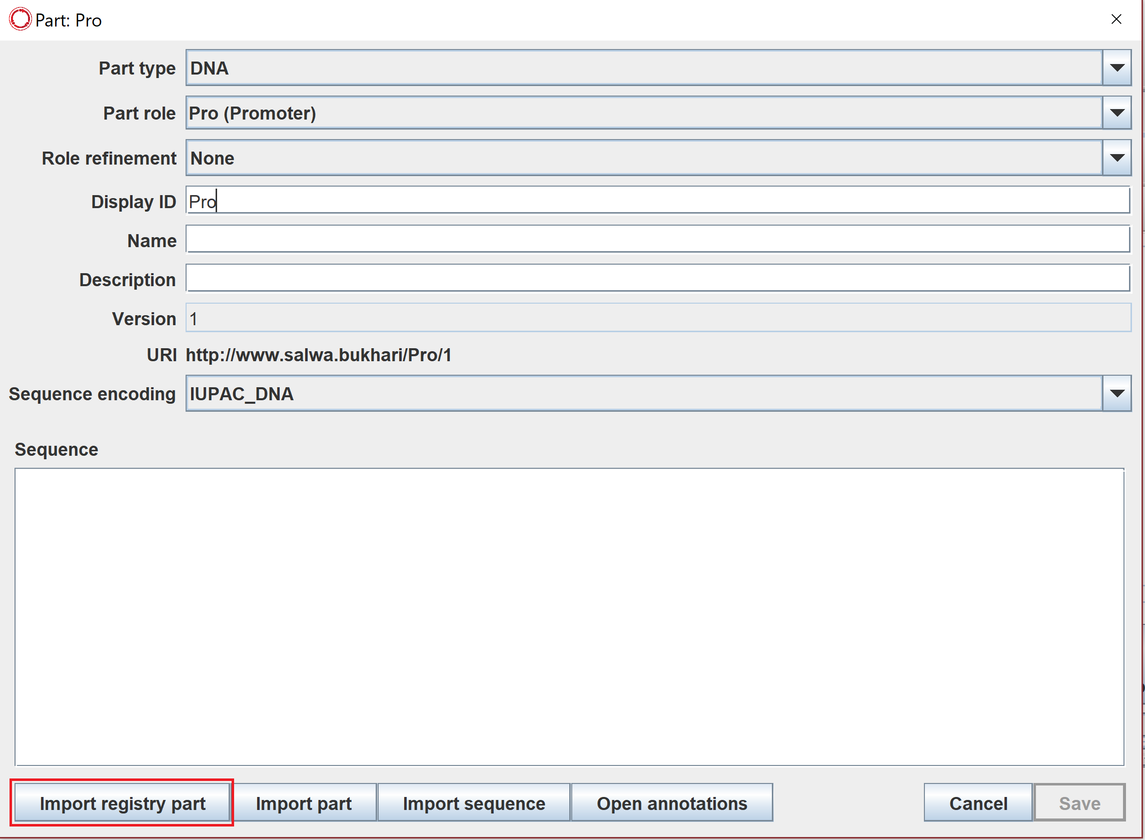
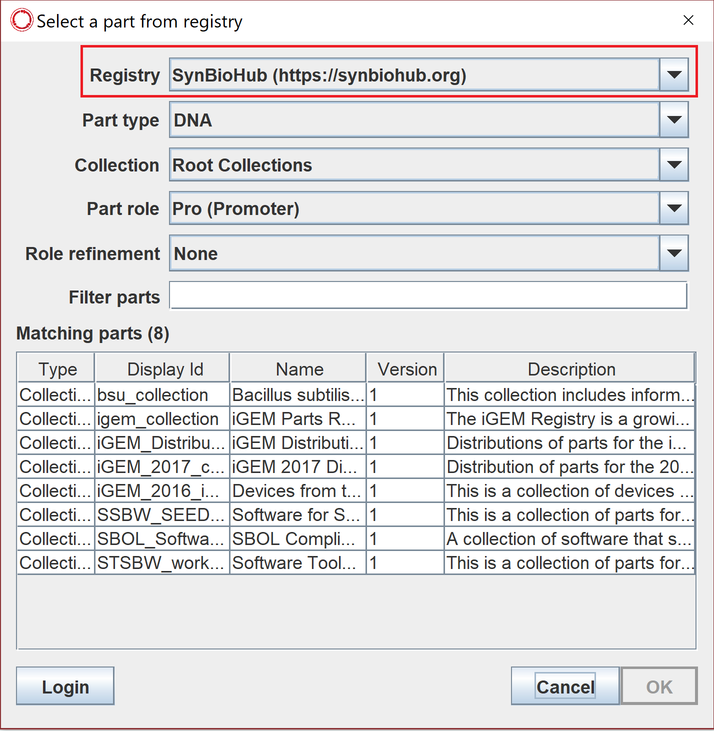
Here is a short video on how to do that.
Uploading to an Online Repository #back to top
You can share your design by uploading it to an online repository such as synBioHub.
First, go to File → upload → Design, as showen below
Second, choose the online repository, make sure you are logged in to synBioHub.
If you need additional help, you can watch this tutorial video on how to upload to SynBioHub.
Exporting to Other File Formats #back to top
iBioSim allows you to export to other file formats such as GenBank or FASTA.
To do that, simply go to Files → export → Design, and then you can choose the desired formats.
Importing from Other File Formats #back to top
iBioSim also allows you to import from other file formats saved on your computer.
Similar to exporting process, go to Files → import → Design/Model, then your desired format.
Design example #back to top
Here is a tutorial video on how to create an example of a full design in iBioSim.
Credits#back to top
The iBioSim tool is being developed by Chris Myers research group at the University of Utah. This project has involved many students including Nathan Barker, Scott Glass, Kevin Jones, Hiroyuki Kuwahara, Curtis Madsen, Nam Nguyen, Tramy Nguyen, Tyler Patterson, Nicholas Roehner, Jason Stevens, Leandro Watanabe, and Zhen Zhang.
To cite iBioSim, please use the latest paper:
Watanabe, Leandro, Tramy Nguyen, Michael Zhang, Zach Zundel, Zhen Zhang, Curtis Madsen, Nicholas Roehner, and Chris Myers. 2018. "IBioSim 3: A Tool for Model-Based Genetic Circuit Design." ACS Synthetic Biology 8 (7): 1560–1563. doi:10.1021/acssynbio.8b00078.
Copyright and license #back to top
IBIOSIM is released under the Apache 2.0 License.
For more information about the Apache license go to apache.org/licenses/.